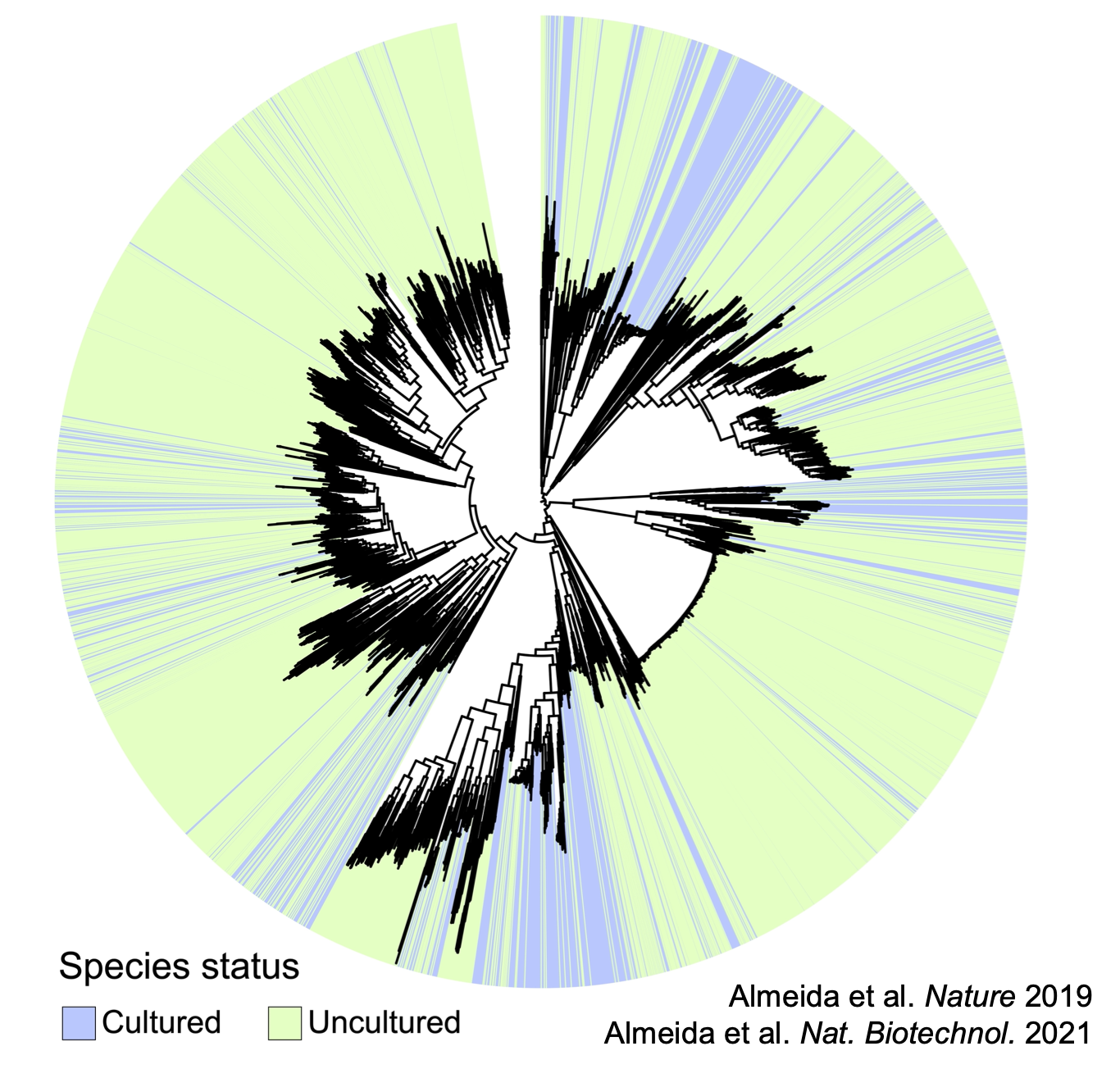
Exploring the uncultured microbiome in health and disease
The study of the human gut microbiome has emerged as a new frontier of biomedical research. With the advent of high-throughput DNA sequencing, a strong link between the human gut microbiome composition and various aspects of human health is being increasingly recognized. However, thousands of bacterial species and viruses in the human gut microbiome are largely unknown, as they remain difficult to culture and experimentally characterize in vitro. This technical limitation represents a major hurdle to understanding the biological mechanisms and beneficial role of the gut microbiome in host health.
Our research group develops and applies large-scale, high-resolution metagenomic approaches to understand the biological role of hidden gut microbes across multiple kingdoms (including bacteria, viruses and archaea) and explore their impact on human health. Some of our key questions are:
- Which uncultured microbiome species are the most clinically relevant?
- What is the role of the gut microbiome in ageing and neurodegeneration?
- How does the gut microbiome influence pathogen colonization and evolution?
- Can viruses and bacteriophages be used as biomarkers of health and disease?
- How does the multi-kingdom microbiome differ across human body sites?
By providing an integrated view of the physiology and functions of the commensal microbiome, we seek to improve our understanding of their role in human health and open new avenues to develop innovative therapeutic applications.
If you are interested in pursuing a PhD or Postdoctoral position in my group feel free to get in touch with a CV and cover letter/statement explaining your interest in our research, so we can discuss future opportunities.
Note: For prospective PhD candidates, we currently have a fully-funded PhD studentship available as part of the Cambridge Biosciences DTP PhD Programme (Targeted studentship, Project: TRG-VET-AA26). Application deadline: December 2nd, 2025. You can find more details here.